Our proprietary functional genomics capabilities, called Foresight, are elucidating disease biology to deliver hyper-targeted treatments to patients with unaddressed tumor mutations. Fore’s experienced team have trained and applied this algorithm for years and are the foremost experts on the functional genomics underlying cancer driver genes, allowing us to identify the right drugs for the right patient populations.
Platform

Most mutations are infrequent, understudied, and unaddressed — even on well-known oncogenes. As such, existing drugs target only the most common cancer-triggering mutations.
Our high-throughput, high-precision functional genomics approach opens up new therapeutic options for patients whose cancers are a result of the multitude of rarer cancer-triggering mutations that remain largely untreatable today.
The better we understand unaddressed mutations, the more patients we can reach. Here’s how.
Strategy
Identify
We identify new mutation targets through functional genomics.
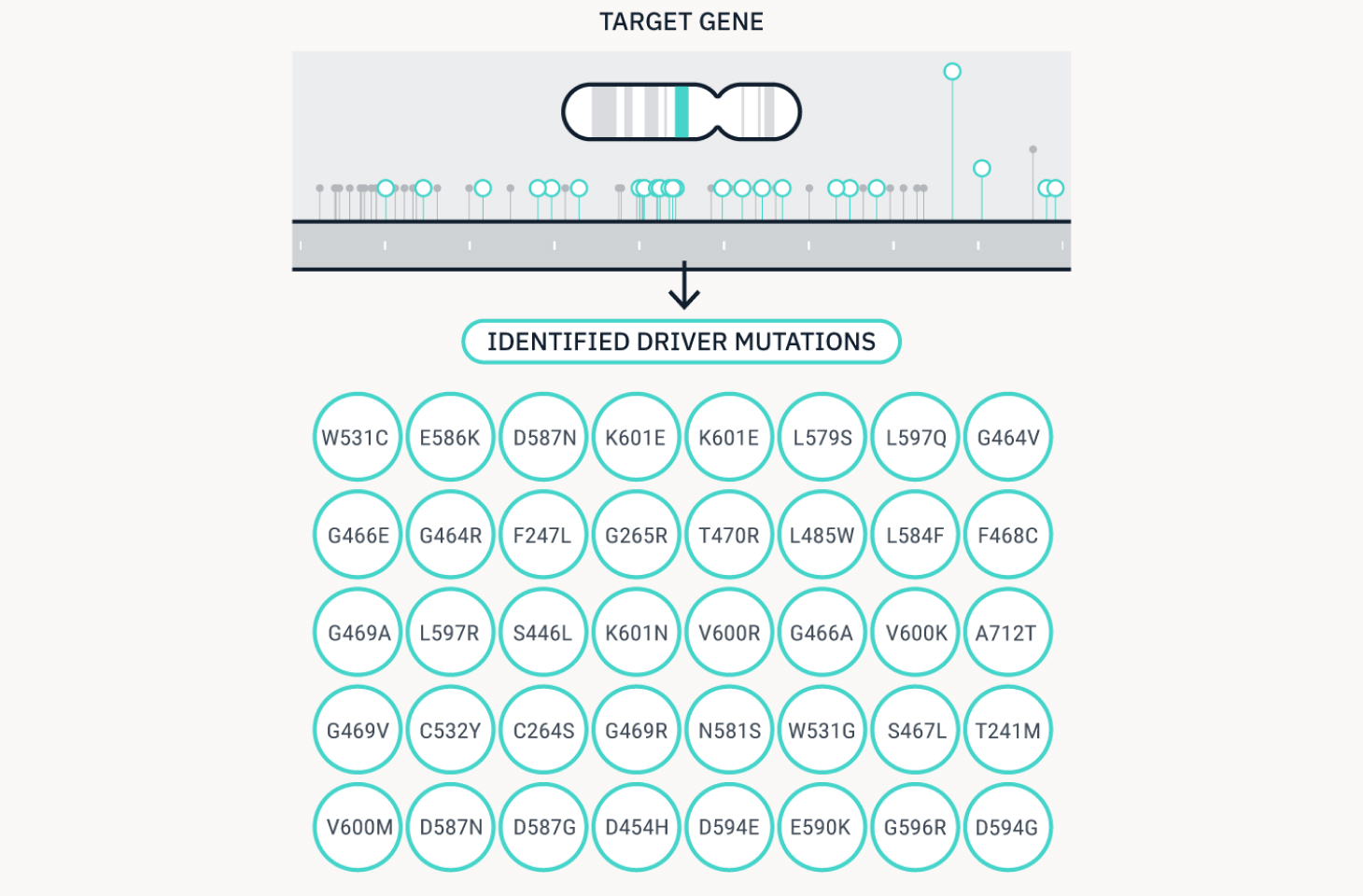
Match
We match hundreds of targets to candidate compounds.
Newly identified driver mutations are retested through Foresight for their response to different compounds. Compounds that reduce oncogenic signaling pathway activity are flagged as potential drug candidates, therefore generating a range of potential new therapeutic applications.
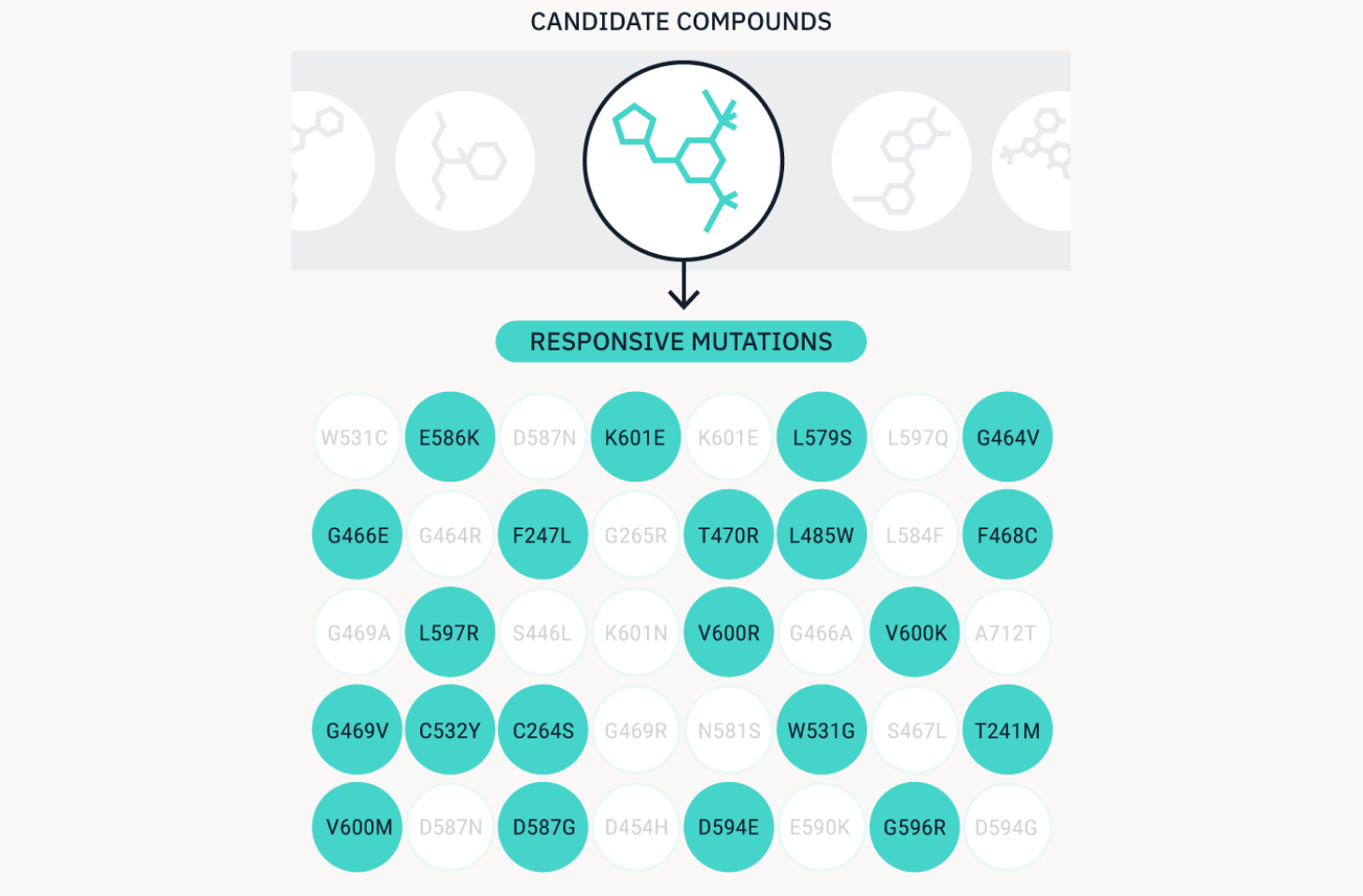
Build
We define new patient subpopulations.
Matching rare oncogenic mutations to theoretically effective drug candidates allows us to pinpoint the diverse patients who are expected to respond to the drug. We can group patients with rare and unaddressed mutations into new populations that have a strong chance of responding to treatment, allowing us to optimize the clinical trial process and accelerate drug development.
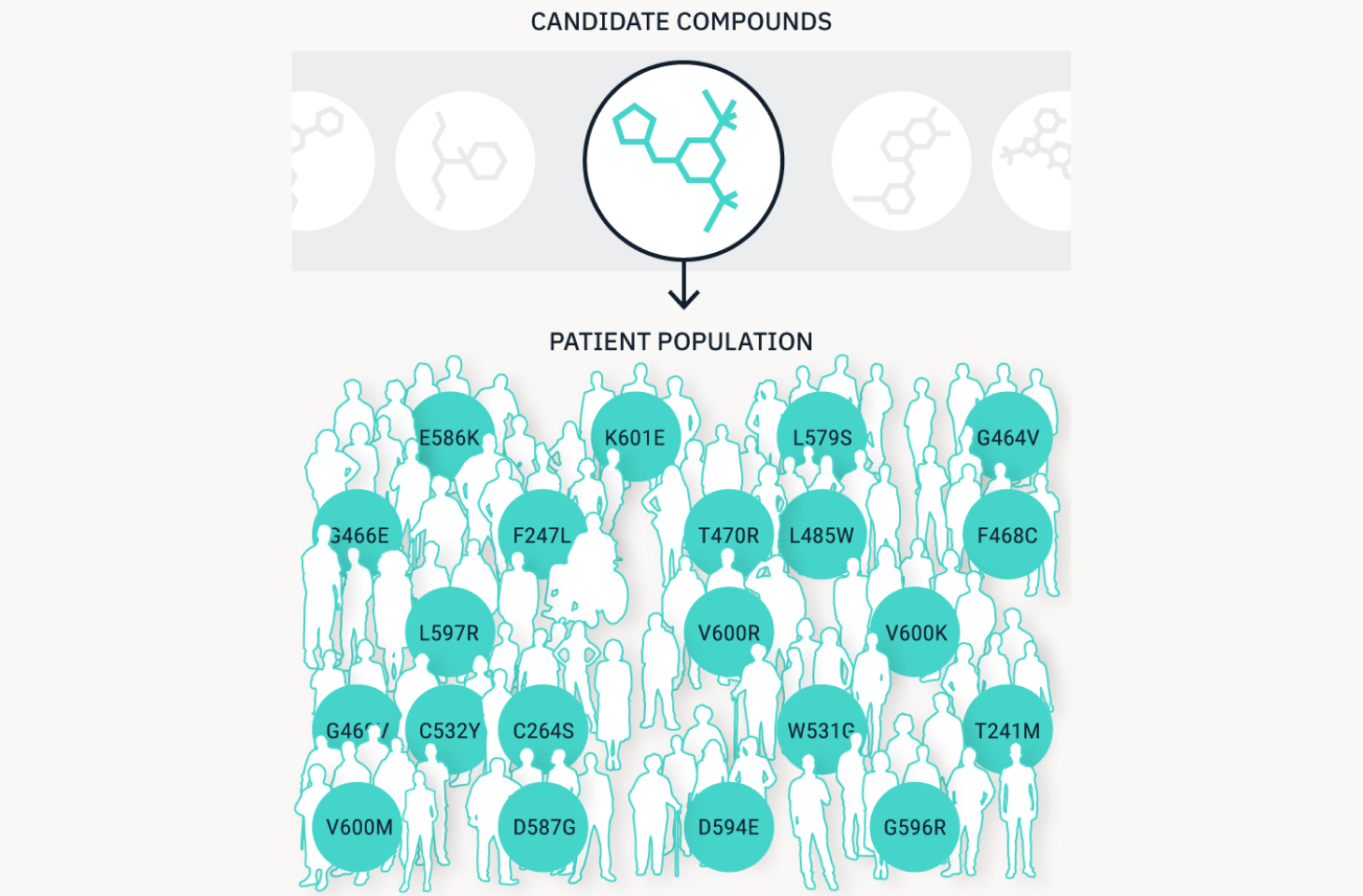
Partner
We put partnerships at the heart of our strategy.
We focus on clinic-ready compounds and can accommodate a wide range of partnering structures. Our approach allows us to determine the best drug candidates without any internal biases, thus circumventing the potential challenges and low probabilities of success associated with early research efforts.
Contact us at oib.e1713552915rof@r1713552915entra1713552915p1713552915 to discuss a potential collaboration.
Foresight Engine
Synthesize
Our unique strategy is supported by novel technologies that merge experimental biology and machine learning at scale.
Our cell models are grounded in human data. We start by identifying addictive targets by using machine learning on population scale data, then use gene editing to synthesize all the mutations reported on our targets.
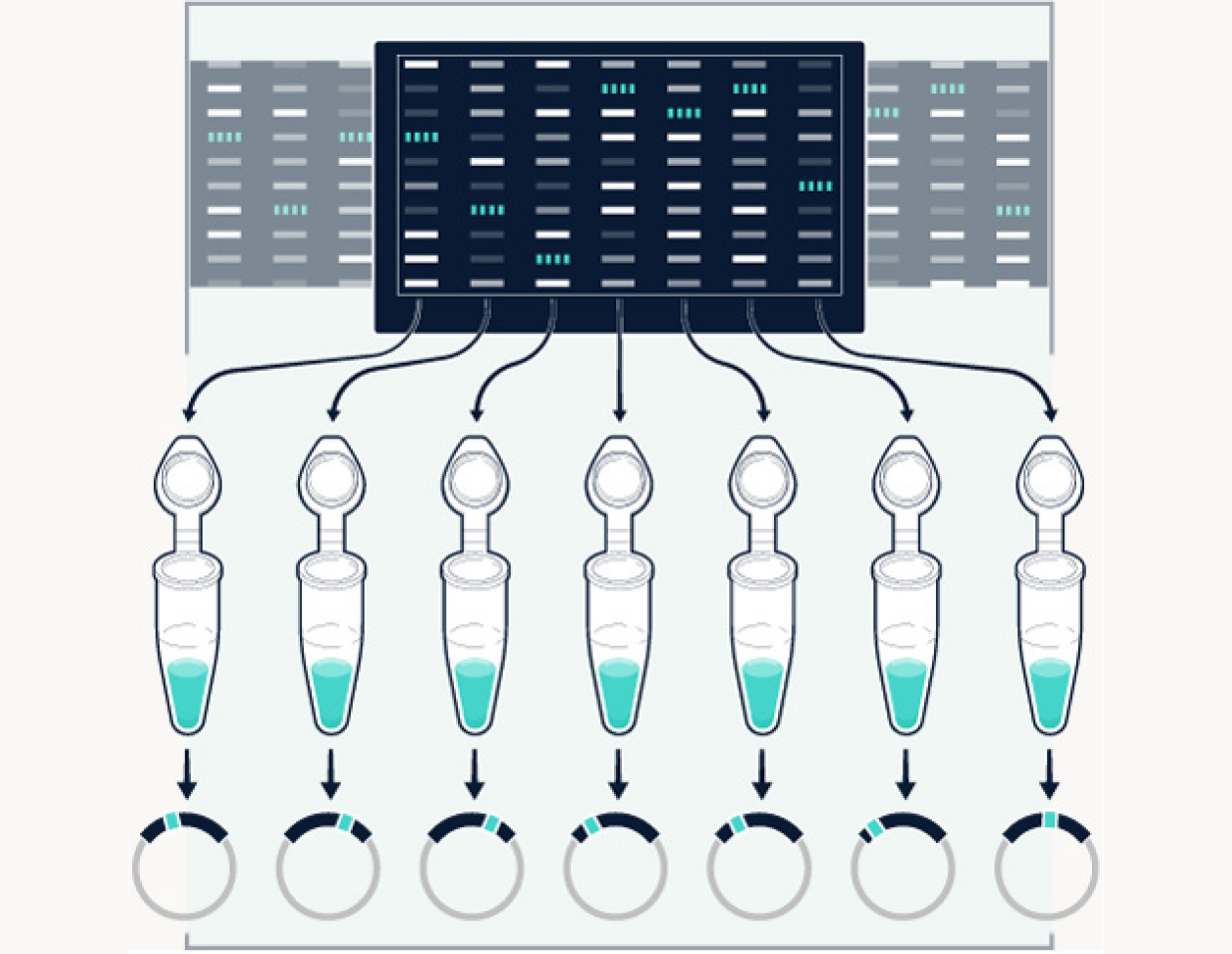
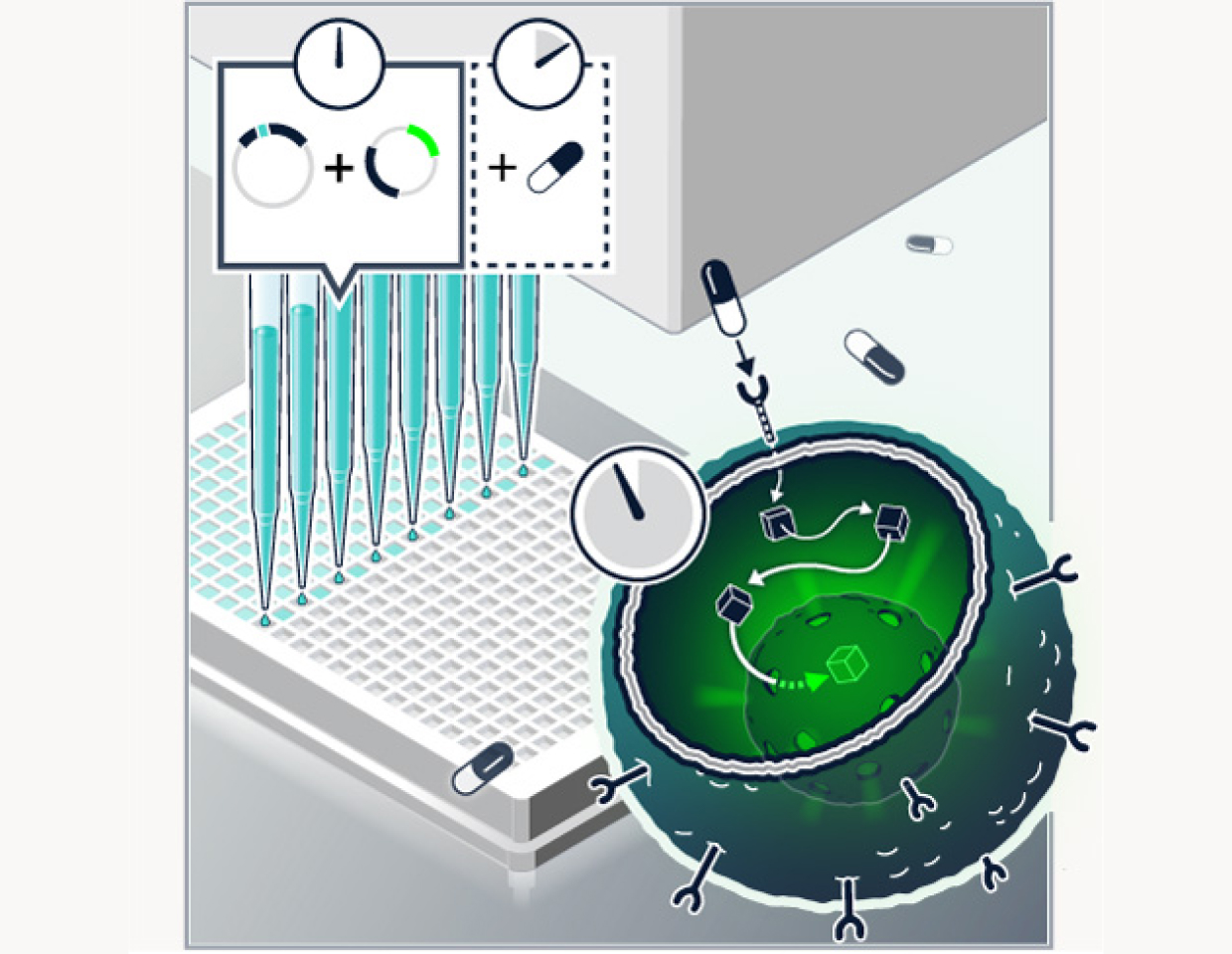
Transfect
The synthesized mutations are transfected into live cells with and without candidate drug compounds, then expressed using a patented method.
Image
Following an incubation period, we deploy high content cellular microscopy to observe the mutations in action, with multiple channels capturing different tagged proteins.
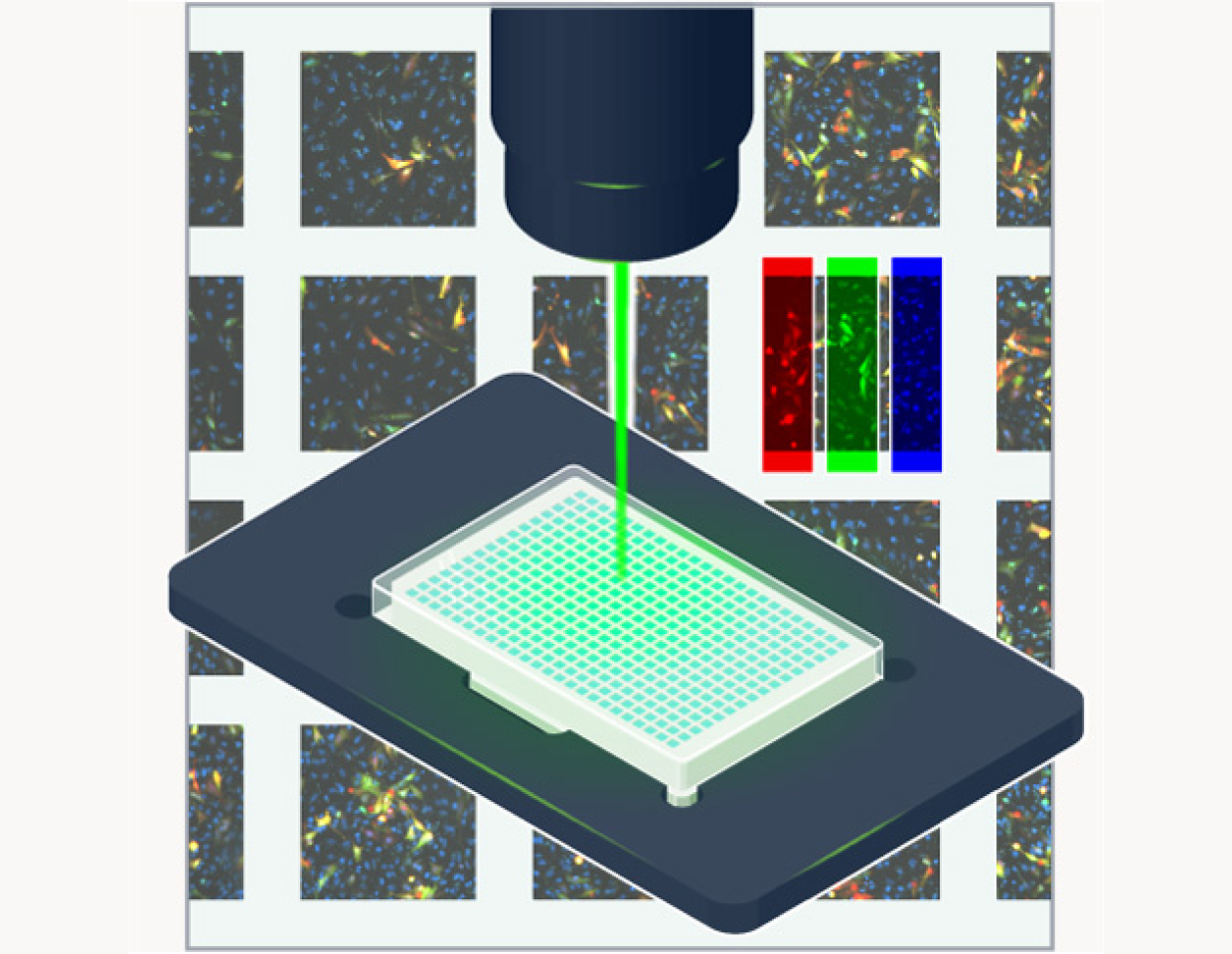

Analyze
We use machine learning to analyze the images and quantify each mutation’s functional effect as well as its response to candidate compounds. We aggregate this information and use it to guide our clinical development programs.
